Bioedit alignment tutorial
- Bioedit alignment tutorial manual#
- Bioedit alignment tutorial full#
- Bioedit alignment tutorial windows#
Split window view for simultaneous and synchronized editing of two different places in the same file - split window vertically or horizontally.Formatted translations of nucleic acid sequences with codon usage summary, choice of one- or three-letter amino acid codes, translation of selected region only of nucleic acid, and choice of start/stop codons.ORF searching with user-defined preferences.Binary file format (BioEdit Project format) for fast open and save of large alignments - the 6205 sequences of the prokaryotic 16S rRNA alignment (29 Mb file) open and save in less than 10 sec on a 233 MHz Pentium.View and manipulate alignments up to 20,000 sequences.View sections of very large matrices with plotter (tested on up to a 5183 x 5183 matrix = 180 Mb file).Matrix plotter and line graphs both have point-and-click data selection and the matrix plotter and 1-D line graphs of matrix data are now dynamically linked RNA comparative analysis, including covariation, potential pairings and mutual information analysis (currently capable of generating matrices up to 10,000 x 10,000 - but this would be a 600+ Mb file) with matrix plotter for 2-D matrix output tables and area graphing for individual rows of a data matrix.Easy customization of menu shortcuts for editor window.Allows import of compatible formats directly from the clipboard without saving to a file first.Utilizes Don Gilbert’s ReadSeq to automatically import and export 11 additional formats, including MSF, ASN.1, IG/Stanford and EMBL.Reads and writes Genbank, Fasta, Phylip 3.2, Phylip 4, and NBRF/PIR formats.Verbally read back sequences in single sequence editor to verify hand-typed sequence entries.Rudimentary phylogenetic tree viewer (for phylip-format trees) that allows node flipping and printing.Append one alignment to the end of another.Merge alignments through a reference sequence.Sort sequences by name, LOCUS, DEFINITION, ACCESSION, PID/NID, REFERENCES, COMMENTS or by residue frequency in a selected column.Specify characters to be considered valid for calculations in amino acid and nucleotide sequences.Lock sequences to prevent accidental edits.
Bioedit alignment tutorial windows#

Also show detail of polylinker and draw moveable arrows and shapes with drawing tools. Easily mark positions, add features with arrows and curved boxes, and mark restriction enzyme cut sites. Plasmid drawing interface for automated creation of plasmid vector graphic from a DNA sequence.
Bioedit alignment tutorial full#
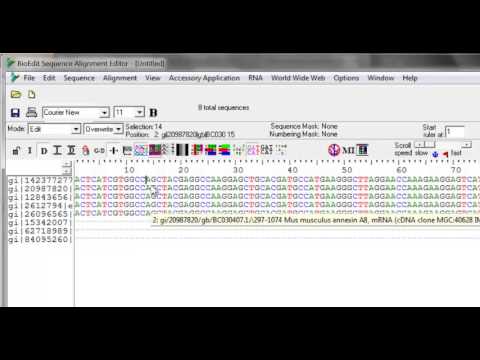
In-color alignment and editing with separate nucleic acid and amino acid color tables and full control over background colors.
